Protein details/Phosphorylation: Difference between revisions
Jump to navigation
Jump to search
No edit summary |
No edit summary |
||
| (4 intermediate revisions by 3 users not shown) | |||
| Line 1: | Line 1: | ||
The Phosphorylation section of the [[Protein | The Phosphorylation section of the [[Protein details]] page in GlyGen provides a detailed list of phosphorylation sites and kinase proteins responsible for phosphorylation. | ||
==Phosphorylation== | ==Phosphorylation== | ||
[[File:Protein Phosphorylation Screenshot.png|thumb|Screenshot of the Phosphorylation section on the [[Protein | [[File:Protein Phosphorylation Screenshot.png|thumb|Screenshot of the Phosphorylation section on the [[Protein details]] page in GlyGen.]] | ||
This section provides the following information: | This section provides the following information: | ||
| Line 14: | Line 14: | ||
The Phosphorylation data is collected and integrated from '''[https://uniprot.org UniProtKB]''', '''[https://pubmed.ncbi.nlm.nih.gov/ PubMed]''', and '''[https://research.bioinformatics.udel.edu/iptmnet/ iPTMnet]''' databases. | The Phosphorylation data is collected and integrated from '''[https://uniprot.org UniProtKB]''', '''[https://pubmed.ncbi.nlm.nih.gov/ PubMed]''', and '''[https://research.bioinformatics.udel.edu/iptmnet/ iPTMnet]''' databases. | ||
*'''[https://www.uniprot.org/ UniProtKB]''' - only reported sites information and predicted information is downloaded from UniProtKB. | |||
*'''[https://research.bioinformatics.udel.edu/iptmnet/ iPTMnet]''' - only reported sites information and predicted information is downloaded from iPTMnet | |||
{{Expand section|small=no}} | {{Expand section|small=no}} | ||
| Line 47: | Line 49: | ||
==Data harmonization== | ==Data harmonization== | ||
{{Expand section|small=no}} | |||
==Data filtering== | ==Data filtering== | ||
{{Expand section|small=no}} | |||
<br /> | <br /> | ||
Latest revision as of 17:39, 9 December 2021
The Phosphorylation section of the Protein details page in GlyGen provides a detailed list of phosphorylation sites and kinase proteins responsible for phosphorylation.
Phosphorylation
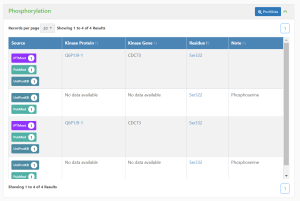
This section provides the following information:
- Source - Evidence linking to the databases and papers that provided the phosphorylation information.
- Kinase Protein - GlyGen ID of the enzyme responsible for catalyzing the covalent addition of a phosphate group to the protein. Eg. Q6P1J9-1
- Kinase Gene - Name of the gene that encodes the kinase protein. Eg. CDC73
- Residue - Residue on which phosphorylation occurs by kinase protein; commonly occurs on amino acids serine and threonine, and occasionally tyrosine. Eg. Ser322
- Notes - Additional information about the entry such as curation notes, phosphorylation subtype, remarks, etc.
Source of information
The Phosphorylation data is collected and integrated from UniProtKB, PubMed, and iPTMnet databases.
- UniProtKB - only reported sites information and predicted information is downloaded from UniProtKB.
- iPTMnet - only reported sites information and predicted information is downloaded from iPTMnet
This section needs expansion. You can help by adding to it. |
Data access
The collected data is processed and stored in data.glygen.org in following datasets.
Homo Sapiens (Human) Datasets
- Human Phosphorylation Sites (UniProtKB; GLY_000274)
- Human Phosphorylation Sites (iPTMNet; GLY_000519)
Hepatitis C Virus Datasets
- HCV1a Phosphorylation Sites (UniProtKB; GLY_000384)
- HCV1b Phosphorylation Sites (UniProtKB; GLY_000385)
SARS Coronavirus Datasets
- SARS-CoV1 Phosphorylation Sites (UniProtKB; GLY_000474)
- SARS-CoV2 Phosphorylation Sites (UniProtKB; GLY_000475)
Mus musculus (Mouse) Datasets
- Mouse Phosphorylation Sites (UniProtKB; GLY_000254)
- Mouse Phosphorylation Sites (iPTMNet; GLY_000520)
Rattus norvegicus (Rat) Datasets
- Rat Phosphorylation Sites (UniProtKB; GLY_000231)
- Rat Phosphorylation Sites (iPTMnet; GLY_000521)
Data harmonization
This section needs expansion. You can help by adding to it. |
Data filtering
This section needs expansion. You can help by adding to it. |