Protein details/Glycosylation: Difference between revisions
| Line 3: | Line 3: | ||
==GlyGen portal== | ==GlyGen portal== | ||
[[File:Protein_Glycosylation_Screenshot.jpg|thumb|upright|Screenshot of the Glycosylation section on the [[Protein information]] page in GlyGen.]] | [[File:Protein_Glycosylation_Screenshot.jpg|thumb|upright|Screenshot of the Glycosylation section on the [[Protein information]] page in GlyGen.]] | ||
Glycosylation is presented in 4 tabs on the [[Protein information]] page in GlyGen. | Glycosylation is presented in 4 tabs on the [[Protein information]] page in GlyGen. The Glycosylation summary provides the overall information about the section like the total number of sites (O,N,S,C linked sites), total number of N linked and O-linked sites with the total number of glycan annotations (structures) observed on the protein. | ||
===Reported Sites with Glycan=== | ===Reported Sites with Glycan=== | ||
| Line 9: | Line 9: | ||
This tab shows a table with the following columns: | This tab shows a table with the following columns: | ||
*'''Source''' - | *'''Source''' - GlyGen evidence linking to the databases and papers that provided the glycosylation information | ||
*'''Type''' - .. | *'''Type''' - Type of glycosylation. Eg. N-linked, O-linked | ||
*'''GlyTouCan ID''' - ... | *'''GlyTouCan ID''' - Unique accession assigned to the registered glycan structure in [https://glytoucan.org/ GlyTouCan] database. Eg. G01543ZX | ||
*'''Glycan Image''' - Image of the glycan | *'''Glycan Image''' - Image of the glycan in SNFG format | ||
*'''Residue''' - | *'''Residue''' - Amino acid residue of the given protein along with its position. | ||
*'''Note''' - | *'''Note''' - Additional information about the entry such as curation notes, O-glycosylation subtype, remarks, etc. | ||
===Reported Sites=== | ===Reported Sites=== | ||
| Line 27: | Line 27: | ||
==Data access== | ==Data access== | ||
{{Expand section|small=no}} | {{Expand section|small=no}} | ||
The Glycosylation data is collected from the resources such as '''[https://www.uniprot.org/ UniProtKB]''', Glyconnect, UniCarbKB, RCSB PDB, The O-GlcNAc Database. The collected data is processed and stored in data.glygen.org in following datasets. | |||
* Human Glycosylation Sites (UniProtKB) | *Human Glycosylation Sites (UniProtKB) | ||
* Mouse Glycosylation Sites (UniProtKB) | *Mouse Glycosylation Sites (UniProtKB) | ||
* Glycosylation Sites - UniCarbKB [Human proteins] | *Glycosylation Sites - UniCarbKB [Human proteins] | ||
* Glycosylation sites - UniCarbKB [Mouse proteins] | *Glycosylation sites - UniCarbKB [Mouse proteins] | ||
* Human Glycosylation Sites (RCSB PDB) | *Human Glycosylation Sites (RCSB PDB) | ||
* Mouse Glycosylation Sites (RCSB PDB) | *Mouse Glycosylation Sites (RCSB PDB) | ||
* Glycosylation sites - UniCarbKB [Rat proteins] | *Glycosylation sites - UniCarbKB [Rat proteins] | ||
* Rat Glycosylation Sites (UniProtKB) | *Rat Glycosylation Sites (UniProtKB) | ||
* Rat Glycosylation Sites (RCSB PDB) | *Rat Glycosylation Sites (RCSB PDB) | ||
* Human Glycosylation Sites (GlyConnect) | *Human Glycosylation Sites (GlyConnect) | ||
* Mouse Glycosylation Sites (GlyConnect) | *Mouse Glycosylation Sites (GlyConnect) | ||
* Rat Glycosylation Sites (GlyConnect) | *Rat Glycosylation Sites (GlyConnect) | ||
* HCV1a Glycosylation Sites (Literature + UniCarbKB) | *HCV1a Glycosylation Sites (Literature + UniCarbKB) | ||
* HCV1a Glycosylation Sites (UniProtKB) | *HCV1a Glycosylation Sites (UniProtKB) | ||
* HCV1b Glycosylation Sites (UniProtKB) | *HCV1b Glycosylation Sites (UniProtKB) | ||
* SARS-CoV2 Glycosylation Sites (UniProtKB) | *SARS-CoV2 Glycosylation Sites (UniProtKB) | ||
* Glycosylation Sites - UniCarbKB [SARS CoV 2 proteins] | *Glycosylation Sites - UniCarbKB [SARS CoV 2 proteins] | ||
* Human Glycosylation Sites [GPTwiki] | *Human Glycosylation Sites [GPTwiki] | ||
* Human Glycosylation Sites [Automatic Literature Mining] [Automatically verified] | *Human Glycosylation Sites [Automatic Literature Mining] [Automatically verified] | ||
* SARS-CoV1 Glycosylation Sites (UniProtKB) | *SARS-CoV1 Glycosylation Sites (UniProtKB) | ||
* Human O-GlcNAc Glycosylation Sites (MCW) | *Human O-GlcNAc Glycosylation Sites (MCW) | ||
* SARS-CoV2 Glycosylation sites | *SARS-CoV2 Glycosylation sites | ||
* Human Glycosylation Sites UniCarbKB Glycomics Study | *Human Glycosylation Sites UniCarbKB Glycomics Study | ||
* SARS-CoV1 Glycosylation Sites (Literature) | *SARS-CoV1 Glycosylation Sites (Literature) | ||
==Data harmonization== | ==Data harmonization== | ||
Revision as of 15:10, 11 November 2021
The glycosylation section of the Protein information page in GlyGen provides the detailed list of glycosylation sites, reported glycans attached to the protein and predicted sites.
GlyGen portal
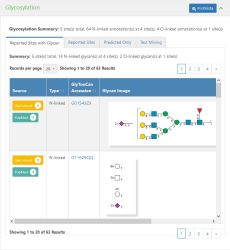
Glycosylation is presented in 4 tabs on the Protein information page in GlyGen. The Glycosylation summary provides the overall information about the section like the total number of sites (O,N,S,C linked sites), total number of N linked and O-linked sites with the total number of glycan annotations (structures) observed on the protein.
Reported Sites with Glycan
This section needs expansion. You can help by adding to it. |
This tab shows a table with the following columns:
- Source - GlyGen evidence linking to the databases and papers that provided the glycosylation information
- Type - Type of glycosylation. Eg. N-linked, O-linked
- GlyTouCan ID - Unique accession assigned to the registered glycan structure in GlyTouCan database. Eg. G01543ZX
- Glycan Image - Image of the glycan in SNFG format
- Residue - Amino acid residue of the given protein along with its position.
- Note - Additional information about the entry such as curation notes, O-glycosylation subtype, remarks, etc.
Reported Sites
This section needs expansion. You can help by adding to it. |
Source of information
This section needs expansion. You can help by adding to it. |
The following resources provided information that was integrated in GlyGen
- UniProtKB - only reported sites information and predicted information is downloaded from UniProtKB
Data access
This section needs expansion. You can help by adding to it. |
The Glycosylation data is collected from the resources such as UniProtKB, Glyconnect, UniCarbKB, RCSB PDB, The O-GlcNAc Database. The collected data is processed and stored in data.glygen.org in following datasets.
- Human Glycosylation Sites (UniProtKB)
- Mouse Glycosylation Sites (UniProtKB)
- Glycosylation Sites - UniCarbKB [Human proteins]
- Glycosylation sites - UniCarbKB [Mouse proteins]
- Human Glycosylation Sites (RCSB PDB)
- Mouse Glycosylation Sites (RCSB PDB)
- Glycosylation sites - UniCarbKB [Rat proteins]
- Rat Glycosylation Sites (UniProtKB)
- Rat Glycosylation Sites (RCSB PDB)
- Human Glycosylation Sites (GlyConnect)
- Mouse Glycosylation Sites (GlyConnect)
- Rat Glycosylation Sites (GlyConnect)
- HCV1a Glycosylation Sites (Literature + UniCarbKB)
- HCV1a Glycosylation Sites (UniProtKB)
- HCV1b Glycosylation Sites (UniProtKB)
- SARS-CoV2 Glycosylation Sites (UniProtKB)
- Glycosylation Sites - UniCarbKB [SARS CoV 2 proteins]
- Human Glycosylation Sites [GPTwiki]
- Human Glycosylation Sites [Automatic Literature Mining] [Automatically verified]
- SARS-CoV1 Glycosylation Sites (UniProtKB)
- Human O-GlcNAc Glycosylation Sites (MCW)
- SARS-CoV2 Glycosylation sites
- Human Glycosylation Sites UniCarbKB Glycomics Study
- SARS-CoV1 Glycosylation Sites (Literature)
Data harmonization
This section needs expansion. You can help by adding to it. |
Data filtering
This section needs expansion. You can help by adding to it. |