SARS-CoV-2 spike glycoprotein
| SARS-CoV-2 spike glycoprotein | |
|---|---|
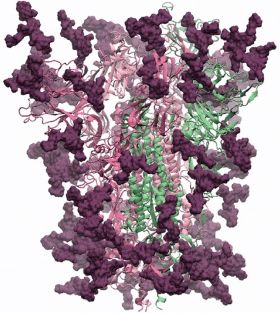 | |
| Glycosylated spike protein | |
| Species: | SARS-CoV-2 |
| NCBI Protein: | QHR63280.2YP_009724390 |
| UniProtKB Protein: | P0DTC2-1 |
The SARS-CoV-2 spike glycoprotein is a large type I transmembrane protein with 1273 amino acids. In addition, this protein is highly glycosylated as it contains up to 18 glycans per protomer. SARS-CoV-2 interacts with the host receptor ACE2 which is also a glycoprotein with several known N-linked glycosylation sites.
Model of the glycoprotein
Initial model (3/30/2020)
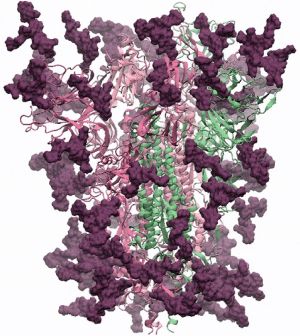
An initial model of the glycoprotein based on the PDB structure 6VSB [1] was created by Prof. Dr. Robert Woods' group from the Complex Carbohydrate Research Center of the University of Georgia. For the GlyGen team Dr. Woods stated:
The glycans (biantennary LacNAc N-glycans) are shown in dark magenta, the protein trimer as ribbons. The protein 3D structure is the Swiss-Model homology model, which is based on PDB ID 6VSB. The glycans were built using GLYCAM-Web. There are 18 glycans per protomer, which comprise ~24% of the mass of the spike protein. Coordinates will be made downloadable as soon as the work is published.
The 3D model is available as:
- Video
Side specific glycosylation model (4/8/2020)
A second model was release from the Woods' group in early April adding specific reported glycans on each glycosylation side of the protein.

The 3D model is available as:
Amino acid sequence
>sp|P0DTC2-1|SPIKE_WCPV Isoform 1 of Spike glycoprotein OS=SARS-CoV-2 OX=2697049 GN=S PE=3 SV=1 CANONICAL=P0DTC2-1 MFVFLVLLPLVSSQCVNLTTRTQLPPAYTNSFTRGVYYPDKVFRSSVLHSTQDLFLPFFS NVTWFHAIHVSGTNGTKRFDNPVLPFNDGVYFASTEKSNIIRGWIFGTTLDSKTQSLLIV NNATNVVIKVCEFQFCNDPFLGVYYHKNNKSWMESEFRVYSSANNCTFEYVSQPFLMDLE GKQGNFKNLREFVFKNIDGYFKIYSKHTPINLVRDLPQGFSALEPLVDLPIGINITRFQT LLALHRSYLTPGDSSSGWTAGAAAYYVGYLQPRTFLLKYNENGTITDAVDCALDPLSETK CTLKSFTVEKGIYQTSNFRVQPTESIVRFPNITNLCPFGEVFNATRFASVYAWNRKRISN CVADYSVLYNSASFSTFKCYGVSPTKLNDLCFTNVYADSFVIRGDEVRQIAPGQTGKIAD YNYKLPDDFTGCVIAWNSNNLDSKVGGNYNYLYRLFRKSNLKPFERDISTEIYQAGSTPC NGVEGFNCYFPLQSYGFQPTNGVGYQPYRVVVLSFELLHAPATVCGPKKSTNLVKNKCVN FNFNGLTGTGVLTESNKKFLPFQQFGRDIADTTDAVRDPQTLEILDITPCSFGGVSVITP GTNTSNQVAVLYQDVNCTEVPVAIHADQLTPTWRVYSTGSNVFQTRAGCLIGAEHVNNSY ECDIPIGAGICASYQTQTNSPRRARSVASQSIIAYTMSLGAENSVAYSNNSIAIPTNFTI SVTTEILPVSMTKTSVDCTMYICGDSTECSNLLLQYGSFCTQLNRALTGIAVEQDKNTQE VFAQVKQIYKTPPIKDFGGFNFSQILPDPSKPSKRSFIEDLLFNKVTLADAGFIKQYGDC LGDIAARDLICAQKFNGLTVLPPLLTDEMIAQYTSALLAGTITSGWTFGAGAALQIPFAM QMAYRFNGIGVTQNVLYENQKLIANQFNSAIGKIQDSLSSTASALGKLQDVVNQNAQALN TLVKQLSSNFGAISSVLNDILSRLDKVEAEVQIDRLITGRLQSLQTYVTQQLIRAAEIRA SANLAATKMSECVLGQSKRVDFCGKGYHLMSFPQSAPHGVVFLHVTYVPAQEKNFTTAPA ICHDGKAHFPREGVFVSNGTHWFVTQRNFYEPQIITTDNTFVSGNCDVVIGIVNNTVYDP LQPELDSFKEELDKYFKNHTSPDVDLGDISGINASVVNIQKEIDRLNEVAKNLNESLIDL QELGKYEQYIKWPWYIWLGFIAGLIAIVMVTIMLCCMTSCCSCLKGCCSCGSCCKFDEDD SEPVLKGVKLHYT
References
- ↑ Wrapp, D.; Wang, N.; Corbett, K. S.; Goldsmith, J. A.; Hsieh, C. L.; Abiona, O.; Graham, B. S.; McLellan, J. S. (2020). "Cryo-EM structure of the 2019-nCoV spike in the prefusion conformation". Science: 1260–1263. doi:10.1126/science.abb2507. PMID 32075877.
External links
- PDB link 6VSB
- https://www.rcsb.org/pdb/explore/remediatedSequence.do?structureId=6VSB
- NCBI Taxonmy
- NCBI Protein